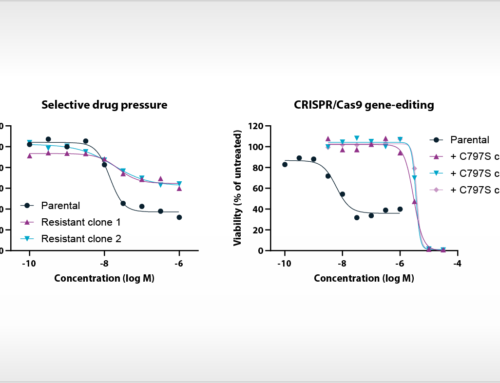
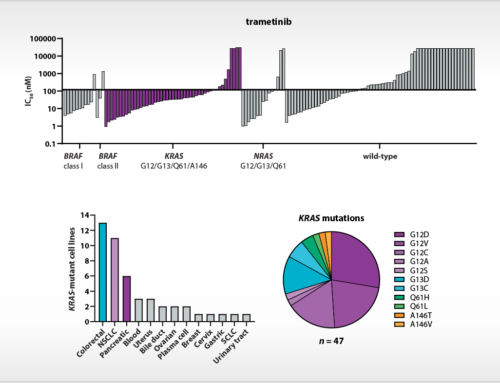
Oss, April 8th, 2016 – Starting from today, NTRC launches a major update of its Oncolines™ bioinformatics platform, referred to as Oncolines™ version 2.0. In the new analysis of cell panel data, 114 new cancer genes are analyzed, including several oncogenes and tumor suppressors that recently have been discovered by massive parallel sequencing of cancer patient genomes, such as ARID1A, MGA, BCLAF1 and others. Importantly, the library of mutations has been validated for their actual presence in patient tumor samples and encompass all genes that are mutated in 5% or more cancer patient samples sequenced in public sequencing projects. The new overview of genetic changes comprises a variety of oncogenic mutations, such as point mutations, frame shifts, insertions, amplifications and deletions. The presence of mutations in the actual Oncolines™ cell line stocks was verified for 23 cancer driver genes, including K-RAS, TP53, CDKN2A, B-RAF, EGFR, CTNNB1, PIK3CA and others, by whole exome or targeted exome sequence analysis.
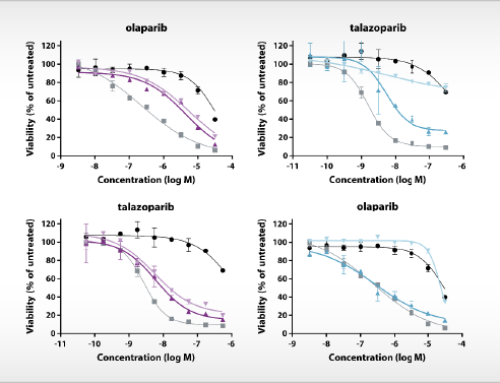
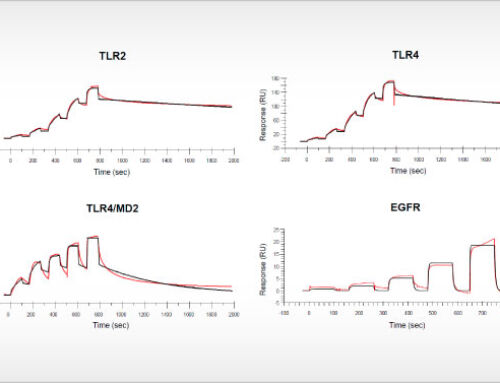
The updated bioinformatic analysis is an integral part of the Oncolines™ compound profiling service. Oncolines™ involves the parallel testing of compounds on 66 human cancer cell line proliferation assays. The analysis has been validated with known targeted therapies [1,2] and enables the development of hypotheses for drug response biomarkers for patient stratification. The genomic biomarker analysis can provide new insight into how drugs or drug candidates interfere with the biology of cancer. By comparative analyses of multiple compounds, Oncolines™ can be used to determine the most precise drug candidate in targeting specific cancer genes, including targeting of difficult-to-drug genes such as K-RAS, MYC and CTNNB1.
At the AACR 2016 meeting in New Orleans, NTRC will present results from the new analysis during the poster session on Wednesday morning. An analysis of the reproducibility of the Oncolines™ profiling of a number of reference inhibitors over a period of more than three years will be presented as well, showing very stable IC50s. In addition to the cell number read-out for Oncolines™ (via ATP content), an apoptosis readout is now available (caspase 3/7). Regular turn-around is four weeks. Further customization of assays is possible.
Presentation of Oncolines™ at AACR 2016, New Orleans
Title: Comparative cancer cell line profiling differentiates the mechanism of action of different PI3K/mTOR, Aurora kinase and EZH2 inhibitors
Presenting author: Joost Uitdehaag
Abstract Number: 2556
Session Category: Experimental and Molecular Therapeutics
Session Title: Cellular Responses to Anticancer Drugs
Session Date and Time: Wednesday Apr 20, 2016 8:00 AM – 12:00 PM
Location: Convention Center, Halls G-J, Poster Section 14
Literature references: [1] PLOS ONE (2014) 9(3) e92146; [2] PLOS ONE (2015) 10(5) e0125021
NTRC is a precision medicine company dedicated to the development of new anti-cancer drugs. NTRC facilitates the development of novel therapies by providing cancer cell line profiling services (Oncolines™, SynergyFinder™ and OncolinesProfiler™) and developing new enabling technologies, such as NFK GreenScreen™. In addition, NTRC develops its own novel targeted therapies based on small molecules, such as selective inhibitors of the TTK (Mps1) protein kinase for chromosomally unstable tumors and inhibitors of the tryptophan metabolizing enzymes IDO1 and TDO for cancer immunotherapy.